Teaching activities developed at Universidad Complutense
Plant of the Day
Description and rubric for students [Spanish, 2022 version]
Description and rubric for students [English, 2019 version]
Plants are somehow familiar to all of us, maybe as a green background, or pretty flowers, but their biology is so different in so many aspects to ours that an extra effort is needed to effectively be able to relate to them as equal living organisms and active agents of their fate. This disconnection between the perception of plants as passive elements of the landscape and their impact in our lifes is often referred to as plant blindness or plant awareness disparity. I use the term “abotanopsia” in Spanish. I have used this concept as a theme for my botany courses ever since 2016, crafting an activity called “The Plant of the Day” [La Planta del Día]. This is a short 5-minute presentation on a particular plant species that students present in pairs towards the middle of every lecture (an opportunity to “change gears” and take a break from a complex topic, for example). The students must choose the species according to their personal interest. Embracing the spirit of Liberal Arts, they are expected to talk, not only about botany, but about connections to other fields (medicine, history, economy, art, etc). This presentation may trigger short discussions on a variety of topics such as the impact of drug dealership in Colombia, a novel cancer therapy, or the role of colonialism in modern crop distributions. This activity is graded with a rubric shared with the students and in general is appreciated as meaningful and enjoyable in their evaluations.
Lab session on Bayesian Inference [Spanish]
Slide show [Spanish]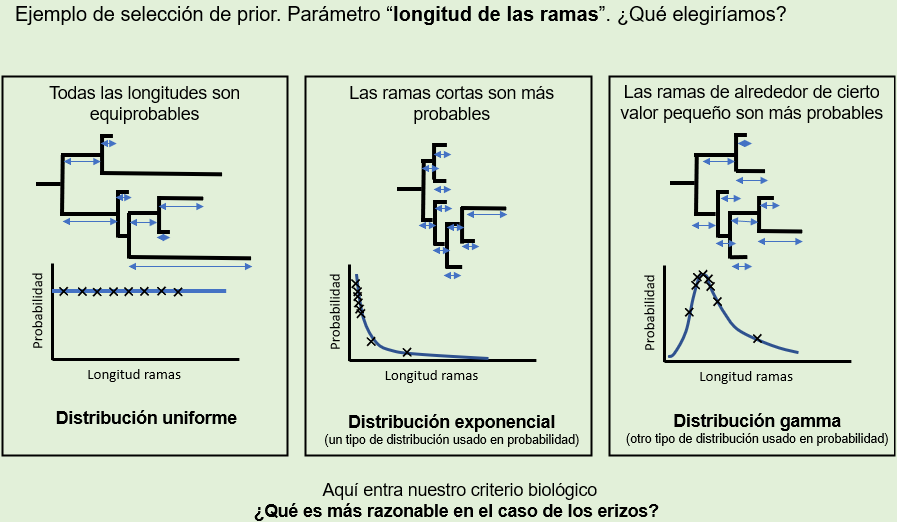
Notes for instructors [Spanish]
2-hour lab session on Bayesian Inference for phylogenetic reconstruction. This session was designed for the Evolutionary Biology course of the Biology Major of the Department of Biodiversity, Ecology, and Evolution of Universidad Complutense de Madrid.
Práctica de dos horas sobre la inferencia bayesiana aplicada a la reconstrucción filogenética diseñada para la asignatura de Biología Evolutiva del Departamento de Biodiversidad, Ecología y Evolución de la UCM.
New courses developed at Augustana College (Illinois)
Techniques in Natural History (BIOL324)
This course was proposed as an introduction to natural history collections in the 21st century (purpose, use, and operation) and it is still taught today in Augie. Students learned how specimens are collected, vouchered, genotyped, and digitized, and as a class they wondered about their role as a tool for the advancement of science and society, including ethical aspects. A significant workload was done in the AUGIE herbarium. Although we used primarily plants, some invertebrates, and slime molds were used as models too. The course combines field work, labs, lectures, and discussions.
Main learning outcomes:
- Sample plants and invertebrates using professional equipment and best ethical practice
- Mount, voucher, digitize, and accession museum/herbarium-quality specimens
- Demonstrate the use of the zoological and botanical nomenclature codes
- Understand the curation process of herbaria and zoological collectiosns
- Perform a basic DNA extraction from a voucher
- Defend, as a professional biologist, the role and contribution of this sort of collections
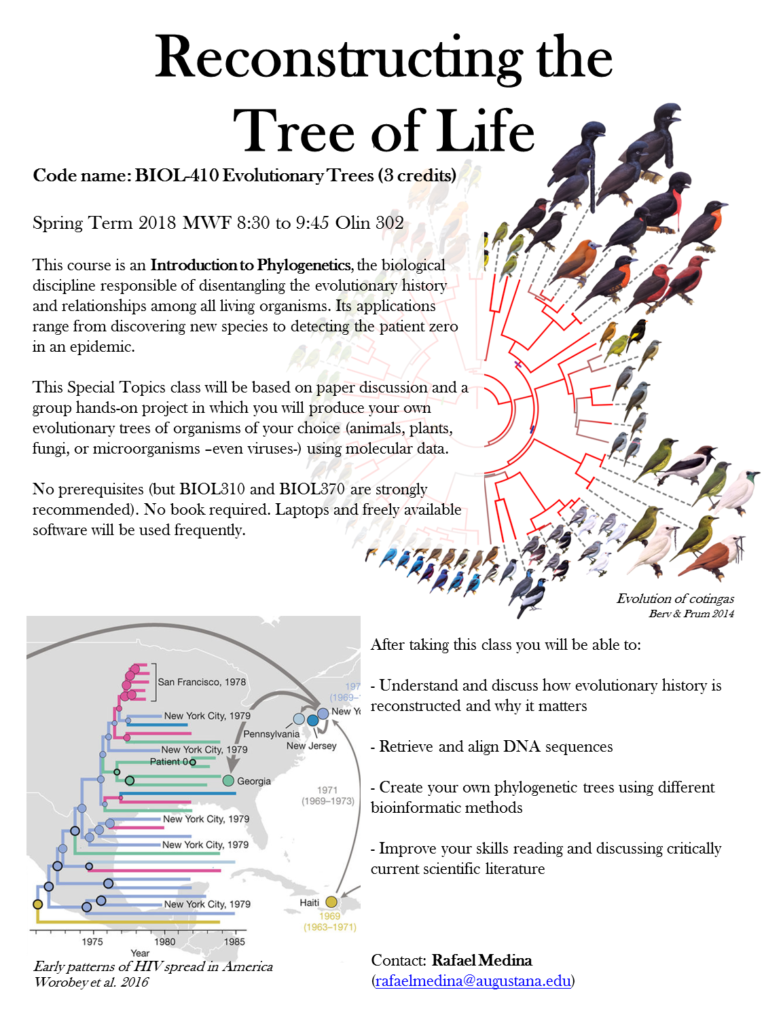
Special Topics: Evolutionary Trees (BIOL410)
This course was an Introduction to Phylogenetics taught once at Augustana College under the trimester calendar. The prompt was that since “Nothing in biology makes sense except in the light of evolution”, students would spend10 weeks making sense out of biology by learning how to see organisms in evolutionary perspective and turn phenomena into stories.
The main learning outcomes were: 1) Understand and discuss how evolutionary history is reconstructed and why it matters; 2) Create your own phylogenetic trees using molecular data freely available and applying different bioinformatic methods; and 3) Improve your skills reading and discussing critically current scientific literature.
Directed Study: Botanical Illustration (BIOL499)
Course Goals: This hands-on course had as its main objective to develop and practice skills and techniques of scientific botanical illustration based both on fresh material and herbarium specimens. The enrolled students received a guided and personalized series of assessments to illustrate plants and reflect on the purpose of this activity. The students deepened into their understanding on plant anatomy and taxonomy. These goals are aligned with the college-wide learning outcomes of Disciplinary Knowledge and Creative Thinking.
Seminar developed at UConn
UCONN EEB3899-007: A primer for practical phylogenetic data gathering
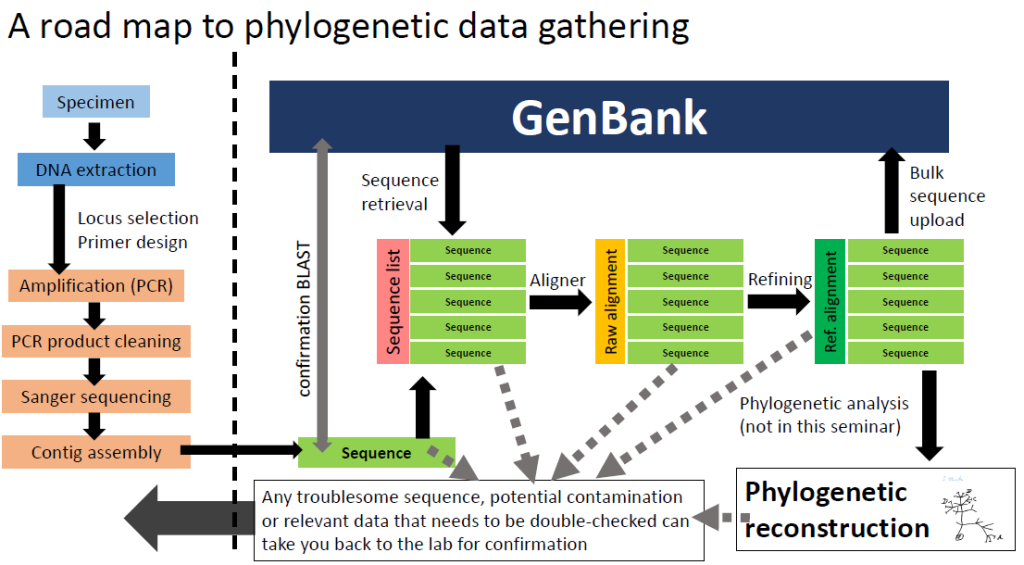 During the Spring semester of 2015, Dr. Yang Liu and I designed and implemented a one-credit seminar specifically targeted for undergraduate students that were starting their experience in research labs as trainees in DNA extraction, amplification and analysis. It is fairly common to see students at the EEB department getting involved in this kind of research, but although they can become competent at the lab tasks, there is usually limited time for them to understand the “big picture” or to receive a proper training in phylogenetic analysis (since these specific skills requires more advanced background and an extensive dedication to be mastered). At the same time, there are many minor skills or activities that they can acquire during their time in the lab (preparation of files for submission to GenBank, primer design, troubleshooting, basic edition of alignments, etc) that we thought should be incorporated into their learning experience.
During the Spring semester of 2015, Dr. Yang Liu and I designed and implemented a one-credit seminar specifically targeted for undergraduate students that were starting their experience in research labs as trainees in DNA extraction, amplification and analysis. It is fairly common to see students at the EEB department getting involved in this kind of research, but although they can become competent at the lab tasks, there is usually limited time for them to understand the “big picture” or to receive a proper training in phylogenetic analysis (since these specific skills requires more advanced background and an extensive dedication to be mastered). At the same time, there are many minor skills or activities that they can acquire during their time in the lab (preparation of files for submission to GenBank, primer design, troubleshooting, basic edition of alignments, etc) that we thought should be incorporated into their learning experience.
We designed the seminar with the following specific objectives, covering first the practical issues that they need to understand for their activity in the lab from the very beginning:
- Background for lab and computer activities: a) Understanding the rationale of the work in a lab (DNA extraction, PCR and Sanger sequencing) as well as providing the tools to develop their own troubleshooting tests; b) A Genbank primer for BLAST, sequence retrieval and preparation of sequence submission and c) Background and basic tools of sequence alignment
- A very brief theoretical introduction to phylogenetic analysis and evolutionary trees
- Discussion of scientific papers or essays dealing with the topics addressed in other sessions
- Generation of distance-based phylogenetic trees
The seminar was structured in 12 sessions of 50 minutes plus a final invited talk:
EEB3899-007_session 1. DNA extraction, amplification and sequencing
EEB3899-007_session 2. Troubleshooting. Nested PCR. Primer design
EEB3899-007_session 3. Sequence manipulation. Retrieving sequences from GenBank
EEB3899-007_session 4. Sequence formats. Alignments
EEB3899-007_session 5. Uploading sequences to GenBank
EEB3899-007 session 6: discussion of the essay “Integrating DNA barcode data and taxonomic practice: Determination, discovery and description” (Goldstein & DeSalle, 2010. Bioessays 33: 135-147)
EEB3899-007_sessions 7 &_8. Evolution, phylogeny and classification. Cladistics. Properties and uses of phylogenetic trees
EEB3899-007 session 9: discussion of the paper “Convergent sequence evolution between echolocating bats and dolphins” (Liu et al., 2015. Current Biology 20: R54)
EEB3899-007_session 10. Using molecules as data. Types of homology. Primary and secondary structure. The ITS region
Session 11: discussion of the paper “Testing reticulation and adaptive convergence in the Grimmiaceae (Bryophyta)” (Henández-Maqueda, Quandt & Muñoz. 2008. Taxon 57: 500-510)
EEB3899-007_session12. Generation of distance-based phylogenetic trees
Final session: an undergraduate student in the Goffinet lab, shared her research experience

